Apurinic/apyrimidinic endonuclease 1 is associated with poor prognosis after curative resection followed by adjuvant chemotherapy in patients with stage III colon cancer
Article information
Abstract
Purpose
Apurinic/apyrimidinic endonuclease 1 (APE1) is a key enzyme involved in the base excision repair pathway. It also has redox activity and maintains various transcription factors in an active reduced state. APE1 may be associated with chemoresistance. In the present study, we first investigated the expression level of APE1 protein and its correlation with oncologic outcomes of oxaliplatin-based chemotherapy in patients with stage III colon cancer. Further, we investigated the effects of human APE1 siRNA on the sensitivity of oxaliplatin in SNU-C2A colon cancer cells.
Methods
Tissue specimens from tumor and normal colon of 33 patients with stage III colon cancer were obtained from 2006 to 2009. The patients received at least eight cycles of oxaliplatin-based chemotherapy. APE1 expression was analyzed by immunohistochemistry and Western blotting using a cultured SNU-C2A cell line. Cell viability and apoptosis were determined by Cell Counting Kit-8 and caspase-3 cleavage using Western blotting.
Results
All the colon cancer tissues showed APE1 staining in the nucleus, whereas all the normal colon tissues were negative for APE1 staining in the cytoplasm. The group with a higher expression of APE1 demonstrated poorer prognosis than the group with low expression (P=0.026 for overall survival and P=0.021 for disease-free survival). Treatment with oxaliplatin resulted in a dose-dependent increase in APE1 expression in SNU-C2A cells. APE1 siRNA significantly enhanced oxaliplatin-induced growth inhibition, and also increased oxaliplatin-induced apoptosis in SNU-C2A cells.
Conclusion
APE1 could be considered a prognostic factor in colon cancer patients treated with oxaliplatin-based chemotherapy.
INTRODUCTION
Colon cancer is the third most common cancer and the fourth leading cause of cancer-related deaths in the world [1]. High-grade dysplasia in colon adenomas serves as an important surrogate marker of colon cancer. Therefore, early detection and removal of adenomas is the most effective strategy to prevent colon cancer [2].
Surgical resection is pivotal in the treatment of colon cancer. However, frequent recurrence is observed even after complete resection of the tumor [3]. Such recurrence may be explained by micrometastases of cancer cells causing extra-colonic spread at the time of resection [4]. Therefore, postoperative adjuvant chemotherapy may play an important role in eradicating micrometastases after surgical resection of colon cancer [5]. Major oncological societies such as American Society of Clinical Oncology (ASCO) and National Comprehensive Cancer Network (NCCN) recommend adjuvant chemotherapy for all patients with stage III colon cancer [6]. Adjuvant chemotherapy for stage II colon cancer is not routinely administered, but patients with high-risk factors for recurrence, such as T4 lesions, poorly differentiated histologic type, inadequately harvested lymph nodes, and presence of bowel perforations could be the appropriate candidates for adjuvant chemotherapy. Chemotherapeutic regimens including fluorouracil, calcium folinate, and oxaliplatin (FOLFOX regimen) are considered to be the gold standard for adjuvant chemotherapy in stage II and III colon cancer in the presence of high-risk factors [7]. However, the recurrence rate after oxaliplatin-based chemotherapy following surgical resection has been reported to be relatively high in these patients [8]. One of the important cause of recurrence may be explained by chemoresistance.
Oxaliplatin is a third-generation of platinum-based drug having in vitro and in vivo activities against colon cancer [9]. An understanding of the biological mechanisms of oxaliplatin action is important to treat chemoresistant colon cancer, although, unfortunately, the molecular mechanisms of this chemoresistance are largely unknown. The molecular target of the platinum-based antineoplastic drugs is DNA. These drugs hinder DNA replication and transcription, resulting in cell death. The possible mechanisms of resistance of platinum-based regimen are decreased intracellular accumulation of platinum, enhanced drug inactivation by metallothionein and glutathione, augmented DNA repair process following DNA damage, and formation of platinum-DNA adducts [10,11]. DNA-repair systems have important roles in protecting the genomic stabilization and integrity by protecting against environmental damage to the DNA. However, augmented DNA repair in the tumor cells may give rise to chemoresistance.
The human apurinic/apyrimidinic endonuclease 1 (APE1, also known as APEX1 or REF1) is one of such enzyme whose expression is associated with poor prognosis in prostate cancer, ovarian cancer, cervical cancer, colon cancer, germ cell tumor, and rhabdomyosarcoma [12]. APE1 is a key enzyme involved in the base excision repair pathway, which plays an important role in DNA repair caused by oxidative and alkylating damage [13]. The apurinic/apyrimidinic (AP) site is mainly repaired by APE1 and accounting for 95% of the total APE1 activity [12]. In addition, it also has redox activities and maintains various transcription factor, such as HIF-1-alpha, p53, NK-κB, CREB, and AP-1, in an active reduced state [12,14]. These two functions may have potential in sensitizing cancer cells to chemotherapeutic agents [15]. We first evaluated the expression of APE1 protein and its correlation with sensitivity and oncologic outcomes in colon cancer patients receiving oxaliplatin-based chemotherapy. Further, we investigated the effects of human APE1 siRNA on the sensitivity of oxaliplatin in SNU-C2A colon cancer cells.
METHODS
Patients and clinical specimens
We reviewed the medical records of the patients who underwent curative resection of colon cancer and received adjuvant chemotherapy in our institute. Among the patients reviewed, a total of 33 patients from July 2006 to June 2009 were included in this study. These patients were treated by curative resection and adjuvant FOLFOX chemotherapy. The data included patients’ characteristics, operative findings, histopathology reports, and follow-up data. Preoperative evaluations consisted of history taking, physical examination, estimation of carcinoembryonic antigen (CEA) level, peripheral blood test, colonoscopy, and computed tomography. The primary colon cancer was characterized by TNM staging, pathological type, and tumor location. The postoperative disease staging was determined according to the criteria defined by the American Joint Committee on Cancer (8th edition). The patients having lymph node involvement (N1 or N2) were included in this study. The present study was reviewed and approved by the Institutional Review Board (IRB) of Chungnam National University Hospital (IRB No. CNUH 201302003) and the informed consent was waived by IRB.
The median age of the patients at the time of operation was 63 years (range, 32–80 years). No chemotherapy or radiotherapy was administered to the patients before surgery of colon cancer. All the patients received >8 cycles of FOLFOX chemotherapy.
Immunohistochemistry and APE1 scoring
The expression of APE1 was evaluated by immunohistochemistry of paraffin-embedded tissue sections obtained from colon cancer and normal colonic mucosa (collected more than 2 cm far from the cancerous lesion). Sections from paraffin blocks having a thickness of 3 μm were used for immunohistochemical staining. Mouse anti-human APE1 monoclonal antibody (Novus, Littleton, CO, USA) was diluted (1:4,000) with a background-reducing diluent (Dako, Carpinteria, CA, USA) and the tissue sections were incubated in the mixture for 30 minutes. All the immunostaining processes were performed using the Ventana Discovery system (Ventana, Tucson, AZ, USA) according to the manufacturer’s instructions. The negative controls were treated identically but without the primary antibody. The entire stained sections of each sample were scanned by a light microscope (×20 or ×40). The slides were reviewed by a pathologist who was unaware of any information related to the enrolled patients. Immunochemical scoring was performed according to the percentage of cell staining and the intensity of staining. Low expression was defined as a weak intensity of staining with positive cell percentage <50% or moderate intensity of staining with positive cell percentage <25%.
Cell line and cell culture
Human colon cancer cell line was used in the present study. SNU-C2A cells were purchased from Korean Cell Line Bank (Seoul, Korea) and cultured in Dulbecco’s modified Eagle’s medium (Thermo Scientific HyClone, Logan, UT, USA) supplemented with 10% (v/v) fetal bovine serum and penicillin (100 U/mL)-streptomycin (100 μg/mL) obtained from GIBCO/BRL Life Technologies (Grand Island, NY, USA). The cells were maintained in a humidified incubator at 37°C in an atmosphere containing 5% CO2 and 95% air.
Cell viability assay
The cells were seeded at 5×103 cells/mL in 96-well microplates and allowed to attach for 24 hours. After drug treatment, cell cytotoxicity and/or proliferation was assessed by Cell Counting Kit-8 (CCK-8; Dojindo Laboratories, Kumamoto, Japan). Briefly, a highly water-soluble tetrazolium salt, WST-8(2-(2-methoxy-4-nitrophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium, monosodium salt), produced an orange-colored water-soluble product called formazan. The amount of formazan dye produced by dehydrogenases present in the cells was directly proportional to the number of living cells. CCK-8 (10 μL) was added to each well and incubated for 3 hours at 37°C. The cell proliferation and cytotoxicity were assessed by estimating the absorbance at 450 nm using a microplate reader. Three replicated wells were used for each experimental condition.
Western blot analysis
The cells were incubated with drugs for 24 hours and washed twice in cold phosphate buffered saline (PBS). The cells were lysed with lysis buffer (10 mM Tris, pH 7.4, 150 mM NaCl, 1 mM ethylenediaminetetraacetic acid, 1% TritonX-100, 0.5% NP-40, 1 mM propidium iodide, 1 mM dithiothreitol, and 1 mM phenylmethylsulfonyl fluoride) and placed on ice for 1 hour with occasional vortexing. It was followed by centrifugation followed at 10,000 rpm for 10 minutes. Each cell lysates (50 μg) was subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis and was transferred on to polyvinylidene difluoride membrane. The blots were blocked with 5% skimmed milk in PBS containing 0.05% Tween-20 for 1 hour at 25°C. The blots were then incubated with primary antibody, followed by incubation with anti-rabbit or anti-mouse horseradish peroxidase-conjugated secondary antibody IgG, and finally visualized with enhanced chemiluminescence.
siRNA transfection
Two siRNAs against APE1 and negative control siRNA were transfected using siRNA Reagent System (Santa Cruz Biotechnology Inc., Santa Cruz, CA, USA) according to the manufacturer’s instructions (siRNA Transfection Protocol). The down-regulation of APE1 gene expression was analyzed by immunofluorescence staining.
Statistical analysis
All statistical analyses were performed using SPSS version 24.0 (IBM Corp., Armonk, NY, USA). The correlation between APE1 expression and clinicopathologic parameters was evaluated using chi-square analysis. A survival curve was plotted using the Kaplan-Meier method, and the log-rank test was used to determine the statistical difference. A forward stepwise selection of covariates was used in the Cox proportional hazard model, and a P-value of <0.1 was defined as the threshold for covariate inclusion. All experiments were repeated three times and are expressed as the mean±standard deviation. Mann-Whitney test was performed and a P-value of <0.05 was considered statistically significant.
RESULTS
APE1 expression in stage III colon cancer patients
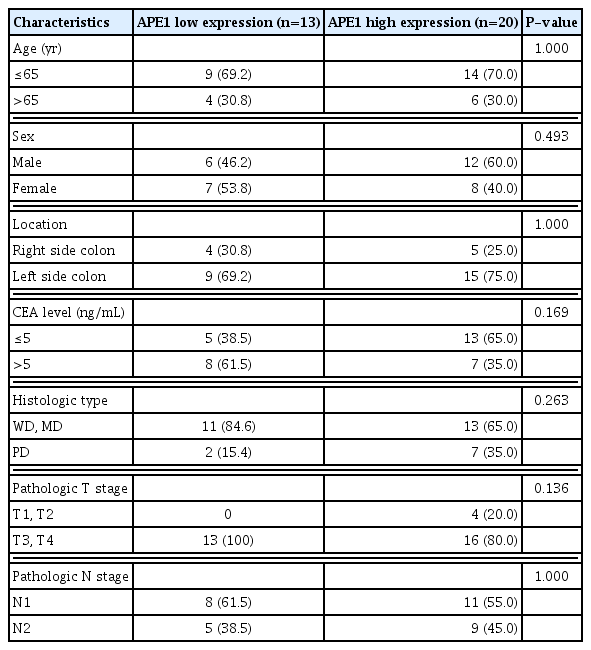
We have investigated the expression of APE1 in 33 colon cancer and normal colon tissue (collected more than 2 cm away from the cancerous lesion) using immunohistochemistry. All the colon cancer tissues showed APE1 staining in the both the nucleus and cytoplasm, whereas all the normal colon tissues showed negative APE1 staining in the cytoplasm. Of the normal colon tissue specimens, 11 specimens (33.3%) showed negative APE1 staining in both the nucleus and cytoplasm (Fig. 1A), while the remaining (66.7%) showed positive APE1 staining in the nucleus (Fig. 1B). Thirty-three percent of the colon cancer tissues demonstrated low APE1 expression (Fig. 1C), and the remaining demonstrated high APE1 expression (Fig. 1D). Clinicopathological parameters were compared according to the findings obtained in immunohistochemical staining (Table 1). The results showed that there were no significant differences between the low and high expression of APE1 expression staining when compared according to age, sex, tumor location, CEA level, and pathological findings.

Immunohistochemical staining for APE1 protein (×100). (A) Negative expression of APE1 in normal colon tissue. (B) High level of APE1 expression in normal colon tissue. Normal colonic mucosa was stained more densely in the nucleus than in the cytoplasm. (C) Low level of expression of APE1 in the adenocarcinoma tissues. The tumor cells were stained more densely in the nucleus than cytoplasm. (D) High level of expression of APE1 in the adenocarcinoma tissues. The tumors cells were densely stained both in the nucleus and cytoplasm. APE1, apurinic/apyrimidinic endonuclease 1.
Correlation between APE1 expression and survival in stage III colon cancer patients
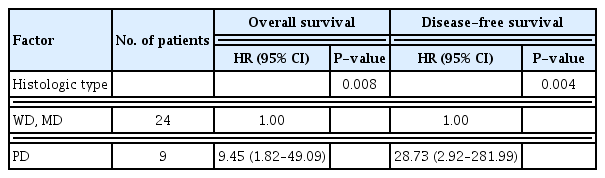
The median follow-up period for all the patients was 51.1 months (range, 13.6–72.7 months) in the present study. During the follow-up period, recurrence occurred in 11 patients (33.3%). The 5-year overall survival and disease-free survival rates were 76.1% and 76.8%, respectively. Table 2 enumerates the findings of univariate analysis showing that different prognostic factors influenced the oncological outcomes. The significant prognostic factors influencing overall and disease-free survival included the histological type of cancer, pathologic N stage, and APE1 expression. In the Kaplan-Meier curve and the log-rank test, the group demonstrating higher APE1 expression was found to have more adverse outcomes in overall survival (P=0.026) and disease-free survival (P=0.021) than the group demonstrating lower APE1 expression (Fig. 2). The variables with P<0.1 from the univariate survival analyses were entered into the Cox proportional hazard model for multivariate analysis of overall and disease-free survival. Poorly differentiated histological type of cancer was an independent poor prognostic factor for both overall survival (P=0.008) and disease-free survival (P=0.004) (Table 3).
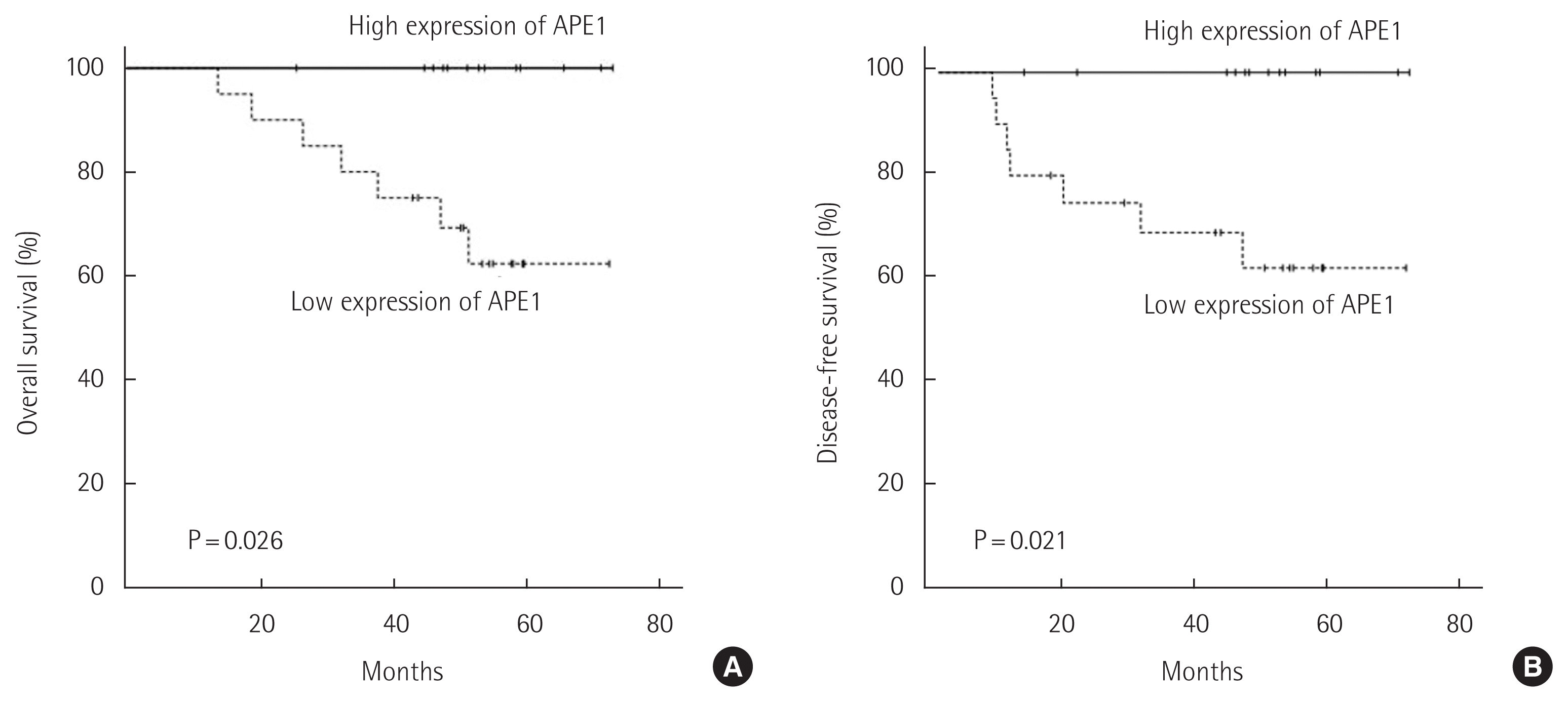
(A) Overall survival and (B) disease-free survival of stage III colon cancer patients who were treated with curative resection and oxaliplatin-based adjuvant chemotherapy according to the results of APE1 immunohistochemical staining. APE1, apurinic/apyrimidinic endonuclease 1.
APE1 expression is induced by oxaliplatin in a dose-dependent manner in SNU-C2A colon cancer cells
By the immunohistochemistry results and survival curve analysis of stage III colon cancer patients, overexpression of APE1 was found to be associated with poor oncological outcomes. From this data, we postulated that APE1 overexpression might be correlated with the resistance of oxaliplatin-based chemotherapy. To determine whether oxaliplatin chemotherapy associated with the expression of APE1, Western blotting was performed with SNU-C2A cells treated with oxaliplatin. APE1 was found to be strongly expressed in SNU-C2A cells treated with ascending doses of oxaliplatin for 24 hours. The higher expression of APE1 with ascending doses of oxaliplatin statistically different (P<0.05 as compared to control) (Fig. 3).

Oxaliplatin-induced APE1 expression in a dose-dependent manner in SNU-C2A cells. The cells were treated with different concentrations of oxaliplatin for 24 hours. Western blot analysis was performed with antibodies for APE1, and GAPDH served as a loading control. APE1, apurinic/apyrimidinic endonuclease 1. a)P<0.05 as compared to control.
APE1 siRNA inhibited oxaliplatin-induced APE1 expression in SNU-C2A cells
To silence the expression of APE1 specifically, SNU-C2A cells were transfected with siRNA-targeting APE1. APE1 expression was determined by confocal microscopy after immunofluorescence staining. As shown in Fig. 4, APE1 siRNA suppressed oxaliplatin-induced APE1 overexpression. This result suggested that APE1 siRNA effectively inhibited APE1 expression in SNU-C2A cells.
APE1 siRNA enhanced the sensitivity of SNU-C2A cells to oxaliplatin
We have determined that APE1 siRNA inhibited APE1 overexpression in oxaliplatin-induced SNU-C2A cells. To determine whether targeting inhibition of APE1 enhanced the activity of oxaliplatin in SNU-C2A cells, the cells were pretreated with APE1 siRNA or control siRNA, and then treated with oxaliplatin at a concentration of 30 μg/mL for 24 hours. The cell viability was determined by CCK-8 assay. Fig. 5 shows a significant decrease in survival of the APE1 siRNA + oxaliplatin treated cells as compared to the control cells treated with siRNA + oxaliplatin. The viability was decreased in 42.1% of SNU-C2A cells treated with APE1 siRNA + oxaliplatin and at 86.6% of SNU-C2A cells treated with control siRNA + oxaliplatin, respectively (P<0.01).

APE1 siRNA enhanced oxaliplatin-induced growth inhibition in SNU-C2A cells. After knockdown with APE1 siRNA, the cells were treated with oxaliplatin (30 μg/mL) for 24 hours. The cell viability was determined by Cell Counting Kit-8 assay. APE1, apurinic/apyrimidinic endonuclease 1. a)P<0.05.
To investigate the effects of APE1 siRNA on apoptosis induction by oxaliplatin in vitro, we estimated the pro-caspase-3/GAPDH ratio by Western blot analysis using caspase-3 antibody (Fig. 6). The pro-caspase-3/GAPDH ratio was slightly increased in SNU-C2A cells treated with oxaliplatin (1.1-fold). However, the pro-caspase-3/GAPDH ratio was significantly decreased in SNU-C2A cells treated with APE1 siRNA+oxaliplatin (0.3-fold, P<0.01 as compared to control), whereas, it was not changed in SNU-C2A cells treated with control siRNA+oxaliplatin (1.0-fold). These results demonstrated that treatment with APE1 siRNA not only enhanced oxaliplatin-mediated growth inhibition but also enhanced oxaliplatin-induced apoptosis in SNU-C2A cells.

APE1 siRNA enhanced oxaliplatin-induced apoptosis in SNU-C2A cells. After knockdown with APE1 siRNA, the cells were treated with oxaliplatin (30 μg/mL) for 24 hours. Apoptosis was measured by the pro-caspase-3/GAPDH ratio with Western blot analysis using caspase-3 antibody. APE1, apurinic/apyrimidinic endonuclease 1. a)P<0.05 as compared to control.
DISCUSSION
APE1 is a ubiquitous and multifunctional protein and its overexpression is often observed in several human tumors [12,16–18]. Limited studies have reported that alteration of APE1 expression is associated with resistance to chemoradiotherapy in head-and-neck cancer and rectal cancer [16,17]. Cisplatin-based chemotherapy in lung cancer was found to be associated with poor oncological outcomes [18]. In the present study, higher expression of APE1 was found to be associated with adverse oncological outcomes in patients with stage III colon cancer receiving oxaliplatin-based adjuvant chemotherapy. APE1 siRNA was found to enhance the sensitivity of SNU-C2A cells to oxaliplatin.
Adjuvant chemotherapy following surgical resection with curative intent in patients with stage III colon cancer can be attributed to a high rate of recurrences. Therefore, authoritative societies such as ASCO and NCCN recommend the use of adjuvant chemotherapy in all patients with stage III colon cancer. An international, multicenter, prospective, randomized clinical trial, conducted by the Multicenter International Study of Oxaliplatin/5-Fluorouracil/Leucovorin in the Adjuvant Treatment of Colon Cancer (MOSAIC) investigators demonstrated that oxaliplatin-based chemotherapy (FOLFOX) improved survival among patients with stage II or III colon cancer [8]. Since reporting of the results of the MOSAIC trial reported, oxaliplatin-based chemotherapy has become the standard adjuvant chemotherapeutic regimen for stage III colon cancer. However, a relatively high rate of recurrence after oxaliplatin-based chemotherapy following surgical resection has been reported in these cases. One of the important causes of this recurrence is chemoresistance. Therefore, chemoresistance remains a significant problem, which physicians need to solve.
Oxaliplatin is a diaminocyclohexane (DACH) derivative of cisplatin. This third-generation of platinum-based drug showed in vitro and in vivo antitumor activities against colorectal cancer [9]. The action of oxaliplatin is thought to be mediated by the retention of the bulky DACH ring during formation of oxaliplatin-DNA adducts, which appear to inhibit DNA replication [10]. DNA repair is pivotal for the removal of oxaliplatin-adducts. Five known DNA repair pathways that protect cellular DNA damage are as follows: (1) nucleotide excision repair, (2) mismatch repair, (3) double-strand break repair, (4) base excision repair, and (5) direct repair [11]. The DNA repair ability of the tumor cells may result in resistance to chemotherapy. Besides the DNA repair pathway, several different mechanisms have been proposed for the tumor cells to become platinum-resistant. However, the molecular mechanisms of oxaliplatin resistance are largely unknown.
APE1 is a key enzyme involved in the base excision repair pathway, which takes part in DNA repair after oxidative and alkylating damage. Therefore, APE1 is important for the protection of tumor cells against the DNA damaging effects of anti-cancer agents. As mentioned, different types of cancers express APE1 [12,16–18]. Kakolyris et al. [19] have previously investigated APE1 expression in colorectal cancer by immunohistochemical staining. They reported predominant APE1 staining in the nuclei of normal mucosa, whereas, only nuclear staining was noted in 11%, and both nuclear and cytoplasmic staining was observed in 39% of colorectal cancer. Kim et al. [17] also studied the expression patterns of APE1 in 83 rectal cancer specimens. Only nuclear staining was noted in 59% and both nuclear and cytoplasmic staining was showed in 37% of the specimens. In this study, all normal colonic mucosa showed negative staining for APE1 in the cytoplasm, and 22 out of the 33 (66.7%) specimens of normal colonic mucosa exhibited APE1 staining (data not shown). In contrast to the normal colonic mucosa, all the colon cancer tissues showed positive staining of both the nucleus and cytoplasm. The reason for discordances in the subcellular localization among different studies might be explained by the variation in the study population. In the present study, only patients with stage III colon cancer were enrolled. Although the subcellular distribution of APE1 varies in normal and tumor cells, several studies demonstrated that cytoplasmic APE1 expression is associated with poor prognosis in different cancers [17,20–22]. In the present study, all the tumor specimens showed cytoplasmic APE1 expression, and a high expression of APE1 in the cytoplasm was found to correlate with more adverse survival outcomes than a low expression of APE1.
The MOSAIC trial demonstrated that 72.2% of the patients with stage III colon cancer after adjuvant FOLFOX chemotherapy had a 3-year disease-free survival [8]. The present study demonstrated similar disease-free survival (76.8%) as compared to the MOSAIC trial. After multivariate survival analysis, poorly differentiated histological subtypes were found to be an independent risk factor affecting overall and disease-free survival in the stage III colon cancer patients who received adjuvant FOLFOX chemotherapy. Although APE1 expression was not an independent factor affecting survival in the multivariate survival analysis, high expression of APE1 was found to be a significant prognostic factor in the univariate survival analysis. A study with a higher sample size could provide further information on the role of APE1 on prognosis in colon cancer overcoming the inherent limitations of this study. Further evaluations are necessary to elucidate the relationship between APE1 and prognosis in colon cancer.
Previous studies have reported that APE1 protein knockdown using siRNA could result in decreased activity of APE1 and increased sensitivity to ionizing radiation, alkylating, and oxidizing DNA damaging agents [23–25]. In the present study, we have demonstrated that APE1 overexpression was associated with the poor oncological outcomes in patients with stage III colon cancer who received FOLFOX adjuvant chemotherapy. The targeted inhibition of APE1 using siRNA was found to enhance the activity of oxaliplatin in the human colon cancer cells. These findings might have both prognostic and therapeutic implications for treating colon cancer.
In addition to DNA base excision repair, APE1 also functions as a redox factor activating various transcription factors, such as HIF-1-alpha, p53, NK-κB, CREB, and AP-1, in an active reduced state [12,14]. These transcription factors are involved in angiogenesis, induction, and progression of cancer cells. Several in vitro studies have demonstrated that inhibition of APE1 decreased the ability of NK-κB to bind to DNA and enhanced the sensitivity of response to chemotherapeutic agents, such as doxorubicin and gemcitabine [26,27]. NK-κB has been linked to chemoresistance in colon cancer cells, and inhibition of NK-κB enhanced the sensitivity of human colon cancer cells to oxaliplatin [28]. Although the redox function of APE1 function has been discovered two decades ago, there is little information on how the redox activity of APE1 influences the response to chemotherapy. The redox function of APE1 controlling the transcription factors contributes to the resistance of the cancer cells to chemotherapy. The relationship between transcription factors and chemotherapy resistance has been increasingly reported [15].
In conclusion, we have reported that the APE1 expression can predict the oncological outcomes and the sensitivity to oxaliplatin-based chemotherapy in stage III colon cancer patients. The in vitro assay demonstrated that the APE1 knockdown could enhance the sensitivity of human colon cancer cells to oxaliplatin. These results suggest that determination of APE1 expression in colon cancer cells before initiating chemotherapy could be a predictor of effective oxaliplatin-based adjuvant chemotherapy in patients with colon cancer.
Notes
CONFLICT OF INTEREST
Jin Soo Kim is an editorial board member of the journal but was not involved in the peer reviewer selection, evaluation, or decision process of this article. No other potential conflicts of interest relevant to this article were reported.